Personal Statement
I have a mixed background of experimental and computational biology, as well as software development.
During my Ph.D., I have generated over 300 Illumina libraries for small RNA-seq, RNA-seq, ChIP-seq, GRO-seq, and Genomic sequencing from genetically engineered Drosophila and mice. In the meanwhile, I developed computational tools for their analyses.
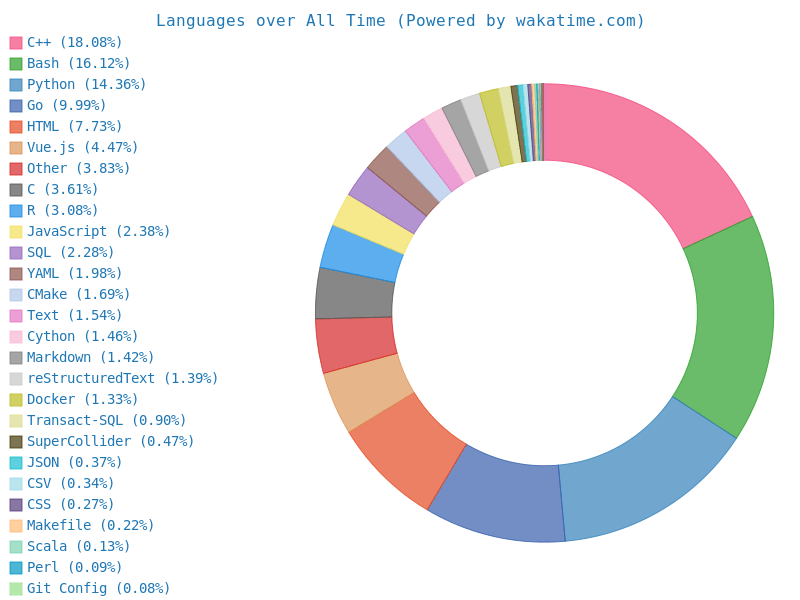
I have more than 10 years of experience using C++ and Python for bioinformatics and algorithm development, and R for statistical calculation, machine learning and figure generation.
I also have experience with web development using Vue.js and Beego, as well as using it for data visualization with D3.js.
Education
Ph.D. in Biomedical Science with Dr. Phillip Zamore
2008/09 - 2015/08
Using Experimental and Computational Strategies to Understand the Biogenesis of microRNAs and piRNAs
In 2008, I joined the Lab of Dr. Zamore to pursue my Ph.D in Biomedical Science to study the biogenesis of small RNAs (including microRNA and piRNA). Since then I have been leading or co-leading 3 experimental-computational hybrid projects—which were published on Science, Molecular Cell, and Current Biology— and 2 computational projects that were published in Nucleic Acids Research and Bioinformatics. The main approach I used in those projects is the combination of next generation sequencing and computational analysis. I have experience analyzing both the secondary generation sequencing (illumina) and the third generation sequencing data (pacBio) using both state-of-the-art tools as well as ad hoc tools developed in my own hands.
Other than my own projects, I constantly provided bioinformatics support for people in my lab and school, some of these works were published in Cell, Molecular Cell, and EMBO Journal.
B.S. in Biological Science
2004/09-2008/05
Job Experiences
Computational Scientist
2016/10-2017/06
Principal Computational Scientist
2017/06-2019/09
Senior Principal Computational Scientist
2019/10-2021/10
Director, Computational Sciences
2021/11-2023/04
Senior Director, Computational Sciences
2023/05-2025/06
VP, Computational Sciences
2025/07-Present
Working closely with genomics team to establish company-wide off-target assessment strategy for CRISPR. Contributed to regulatory submissions including INDs, RTQs, and BLA preparations, providing computational evidence packages and data interpretation to support clinical development programs.
Developed pipelines for various NGS (e.g., Amplicon-, RNA-, scRNA-, WGS/WES-, SITE-Seq) sequencing analysis, mostly with algorithms implemented in C/C++, data visualization in R (rmarkdown, ggplot2, plotly, shiny) and Javascript (D3), as well as Python and Bash.
Implemented an architecture to automatically execute pipelines (upon sequencing finishes or user triggers) using Vue.js, plain Go, MySQL, AWS parallel cluster, SGE, AWS SDK, and Docker.
Senior Bioinformatics Scientist
2015-2016
Developed a Hidden Markov Model based PolyA trimming program with C++
Wrote a pipeline to combine illumina and PacBio data for transcriptome annotation and quantification
Technical skills
C++ 14 years
Python 12 years
R 11 years
Bash 12 years
Javascript 8 years
HTML/CSS 8 years
MySQL 8 years
Go 5 year
Software
piPipes
A set of pipelines for piRNA and transposon analysis via small RNA-seq, RNA-seq, degradome- and CAGE-seq, ChIP-seq, and genomic DNA sequencing
Tailor
A Burrow-Wheeler Transform based short reads aligner specialized in capturing non-templated nucleotide addition to the 3′ ends of small RNAs
IsoSeq PolyA Trimmer
A Hidden Markov model based PolyA trimmer for third-generation sequencing data
Misc Tools
Web-based small utilities